New Docking Software for Virtual Screening and Combinatorial Library Design
Press Release Summary:
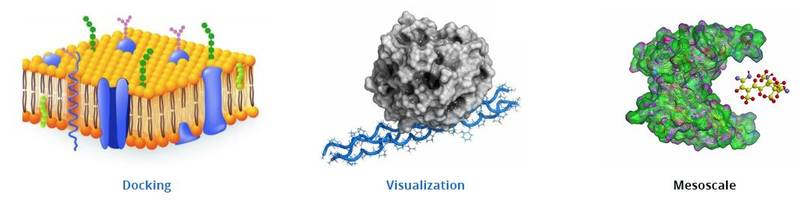
- Discovery Studio is for modeling small and polymeric systems
- Surflex-Dock Molecular docking aims to search for the most suitable way that binds small molecule ligands to receptor molecules
- rDock is designed for high-throughput virtual screening and prediction of binding modes
Original Press Release:
CD ComputaBio Introduces Four Docking Software for Drug Development
CD ComputaBio, a reliable computational service provider in the field of biology, is committed to assisting research and trials, as well as accessing the latest software, technologies, and expertise at a competitive price and fast turnaround for researchers. Molecular docking is a popular technique in drug design for predicting both binding patterns and binding affinity. CD ComputaBio has introduced four molecular docking software: AutoDock, Surflex-Dock, rDock, and Discovery Studio, to help your virtual screening and drug development projects.
The essence of molecular docking is a recognition process between two or more molecules that involve spatial and energetic matching. The docking software places the ligand/small molecule at the active site of the receptor target and by continuously optimizing the position, conformation, dihedral angle of the rotatable bond, as well as the side chain and backbone of the amino acid residues to search for the optimal conformation that binds the ligand/small molecule to the receptor target, thus predicting its binding mode and affinity.
AutoDock
AutoDock is a molecular modeling simulation software particularly effective for protein-ligand docking. AutoDock is one of the most cited docking software applications in the research community. Its applications include but are not limited to X-ray crystallography, structure-based drug design, lead compound optimization, virtual screening, combinatorial library design, and protein-protein docking.
Surflex-Dock
Molecular docking aims to search for the most suitable way that binds small molecule ligands to receptor molecules. Therefore, with the support of the Surflex-Dock software, CD ComputaBio strives to solve the most critical issue confronted by molecular docking, hence, discovering the best binding sites and evaluating related binding strengths between molecules.
rDock
rDock is an open-source molecular docking software designed primarily for high-throughput virtual screening and prediction of binding modes. rDock was developed by Vernalis for high-throughput VS (HTVS) applications. Evolving from RiboDock, this program can be used for proteins and nucleic acids and is designed to be very computationally efficient and allow users to incorporate additional constraints and information as biases to guide docking.
Discovery Studio
Discovery Studio (DS) is a suite of software for modeling small and polymeric systems. On the DS software platform, CD ComputaBio can provide customers with a full range of biopharmaceutical designs and perform hybrid QM/MM calculation, pharmacophore modeling, polymer design and validation, and more.
“Computational science provides essential tools for the next-generation biological science efforts, from managing large volumes of data and centralizing the direction of experimental research to providing knowledge and insight that is not yet available in biological phenomena. CD ComputaBio provides services and software based on cutting-edge computational platforms and technologies with accuracy and sensitivity,” the product manager commented.
About CD ComputaBio
With years of experience, CD ComputaBio can provide customers with professional computational biology services. Utilizing rich experience and powerful technology in computational science, the company can provide customers with assorted computational biology analysis services, such as molecular dynamics simulation, drug design, virtual screening, quantum chemical calculation, etc.
Media Contact:
Company Name: CD ComputaBio
Contact Person: Vivian Smith
Phone: +1-631-371-4691
Country: United States
Website: www.computabio.com




