NIST Method facilitates pnuemonia vaccine testing.
Share:
Press Release Summary:

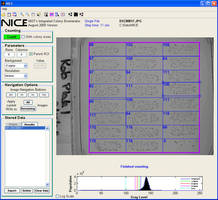
NIST scientists improved efficiency of a pneumonia vaccine testing method developed at the University of Alabama at Birmingham (UAB). The open-source software, NIST's Integrated Colony Enumerator (NICE), takes a digital image that has been loaded into a computer and counts colonies grown from single pneumococcal cells. Windows®-based solution works on images from digital cameras and flatbed scanners, obtaining results with only mean difference of 3% from manual counting method.
Original Press Release:
Scientists Create NICE Solution to Pneumonia Vaccine Testing Problems
Medical clinics the world over could benefit from new software* created at the National Institute of Standards and Technology (NIST), where a team of scientists has found a way to improve the efficiency of a pneumonia vaccine testing method developed at the University of Alabama at Birmingham (UAB).
Pneumonia is the world's leading cause of death in children under five years of age and poses a serious risk to elderly adults. The leading cause of pneumonia worldwide is the pneumococcus bacterium, which also causes meningitis, sepsis and other complications. Pneumococcus has more than 90 strains that vary by geographic region and change over time. Consequently, ongoing testing is necessary to monitor existing vaccines and advance new ones.
One novel, high-throughput testing method involves culturing the bacteria along with a vaccinated person's blood serum and human white blood cells. If the vaccine is effective, the white cells kill the pneumococci and very few of the bacteria survive. Scientists can determine the vaccine's effectiveness by counting the number of surviving pneumococcus colonies, so rapid, accurate and standardized counting of these colonies is critical to this testing method.
At present, the most commonly used counting process is manual counting, which is both time-consuming and exhausting. "Automated counting devices do exist, but they require customized image acquisition methods, are very expensive and are not accessible in impoverished or developing regions of the world. These limiting factors can mean the difference between life and death," says NIST biophysicist Jeeseong Hwang. "So we created software that can be tweaked to work on any common imaging device."
The open-source software, called NIST's Integrated Colony Enumerator (NICE), takes a digital image that has been loaded into a computer and counts colonies grown from single pneumococcal cells. The Microsoft Windows-based software works on images from both digital cameras and flatbed scanners, which are widely available and inexpensive, costing less than $1,000 each. "NICE obtains results that agree well with manual counting, the current gold standard," says Matthew Clarke, who developed the program algorithm and code in Hwang's group. "There's a mean difference of only 3 percent between the two methods."
The project grew from informal talks between NIST and UAB, where researcher Moon H. Nahm developed the testing method. Nahm has been working with vaccine testing methods to support projects by the National Institutes of Health and PATH, a nonprofit organization dedicated to improving health in poor communities around the world. As NIST scientists have been developing ways to standardize counting particles, PATH provided funding and worked with NIST. After 18 months of effort, NICE is now ready.
"We have already identified several clinics in Asia, where we feel many of the potential users are," Hwang says. "We're hoping to get feedback from them so we can improve the software in the future."
* The two software files required to run NICE, along with additional images demonstrating their use, can be downloaded at ftp://ftp.nist.gov/pub/physics/mlclarke/NICE. The files, written in the language MATLAB R2008B, are MCRInstaller.exe (MATLAB compiler engine 7.9) and NICE_B1r.exe (launches NICE). Further information is available from the programmer at nice@nist.gov.
Media Contact: Chad Boutin, boutin@nist.gov, (301) 975-4261




